All topics


Outline how the Hardy-Weinberg equation is derived, including the assumptions for its use.

Which sex chromosome(s) can be found in a human ovum?

Researchers often use short pieces of DNA called Alu sequences to study relationships within populations and between species. The function of these sequences is not known. However, they have the ability to copy themselves at random and insert into other parts of the genome, increasing the DNA fragment size. Alu sequences have inserted into the genome of primates throughout their evolution. One particular group of sequences (Ye) began inserting into the genome relatively early in the evolution of Hominidae and continues to insert itself at low rates. The gel electrophoresis samples below were used to determine the phylogenetic origin of individual Ye5 sequences in primates. The larger DNA fragment in each case contains the Ye5 insert and the smaller fragments are the same region of DNA but without the insert. The green monkey and the owl monkey diverged from the Hominidae approximately 25 million years ago. 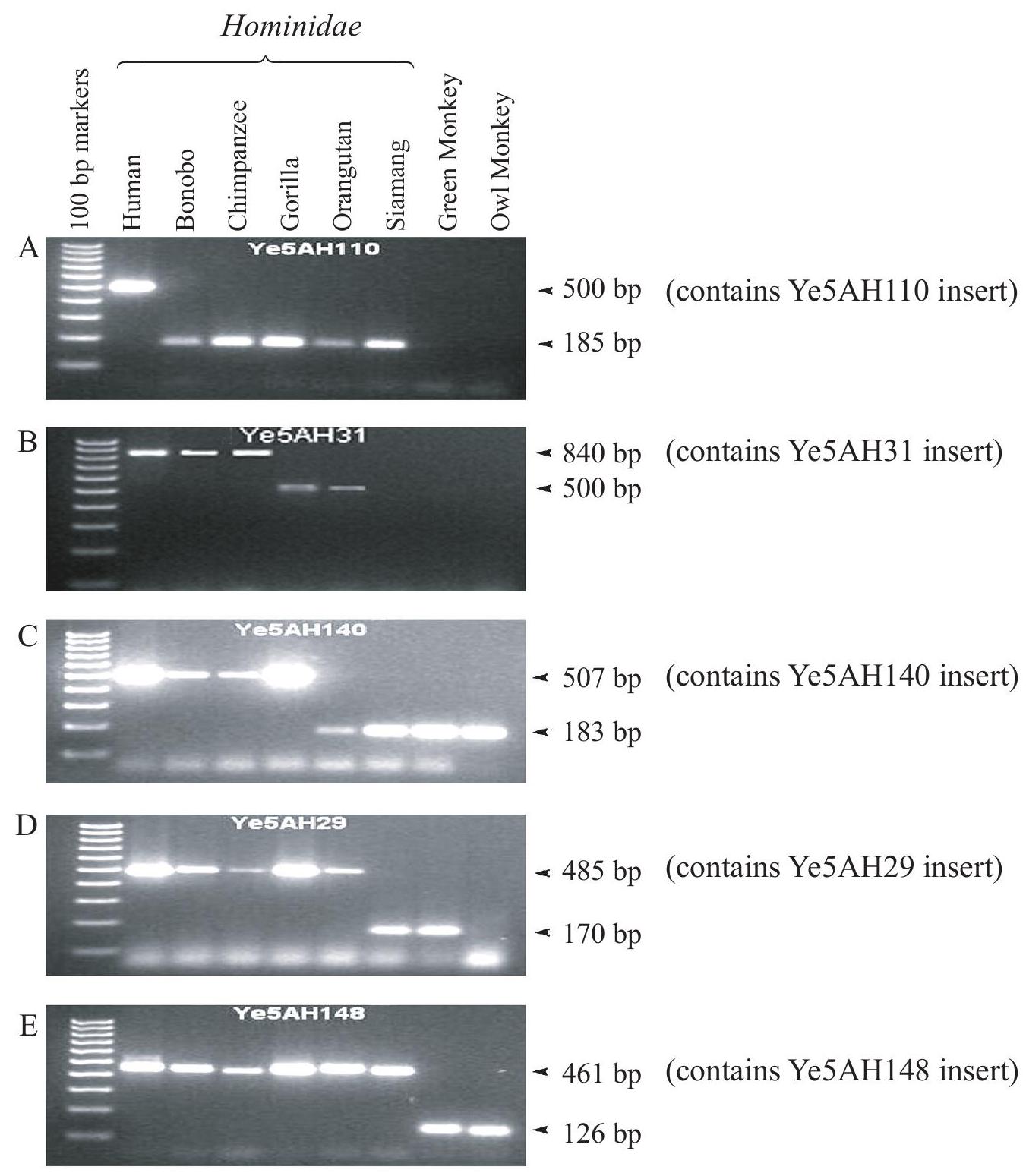

State which Alu insert is found in all of the Hominidae species but not in the other species.

Calculate the size of the Ye5AH140 insert.

Identify, giving a reason, which insert has the most recent origin.

Analyse the data to determine whether bonobos are most closely related to chimpanzees or orangutans.

What is a difference between the features of monocotyledons and dicotyledons?
| Monocotyledons | Dicotyledons | |
|---|---|---|
| A. | Tap roots | Fibrous roots |
| B. | Two seed-leaves | One seed-leaf |
| C. | Vascular tissue in rings in the stem | Vascular tissue scattered through the stem |
| D. | Parallel-veined leaves | Net-veined leaves |

There is evidence that humans, chimpanzees and gorillas evolved from a common ancestor. Biologists recently selected a large number of genes at random from the human genome and compared their base sequences with those of the same genes in chimpanzees and gorillas. The number of base substitutions in the evolution of each species from the common ancestor was then deduced. Some of the base substitutions did not cause a change in the amino acid sequence encoded by the gene. These mutations were assumed to be neutral in terms of natural selection and were used to estimate the total mutation rate in each species. The other base substitutions did cause a change in the amino acid sequence encoded by the gene, each base substitution causing one amino acid substitution. The estimated mutation rate was then used to calculate an expected number of amino acid substitutions. The results are shown in the table below. | Species | Mutation rate per nucleotide per year | Observed number of amino acid substitutions per generation | Expected number of amino acid substitutions per generation | | :--- | :---: | :---: | :---: | | Human | 0.00000000133 | 2.6 | 4.2 | | Chimpanzee | 0.00000000122 | 1.5 | 3.2 | | Gorilla | 0.00000000123 | 1.9 | 3.1 | (Source: Crow, Nature (1999), 397 pages 344-347)

Identify which of the three primate species has evolved most rapidly since divergence, with a reason based on the data in the table.

Analyse the data in the table to find in which species most mutations have been eliminated per generation. Show clearly how you have reached your conclusion.

Using the data in the table, evaluate the stability of DNA as a genetic material.

What is a difference between two alleles of a gene?

Which disease is an example of sex-linked (X-linked) inheritance?

Palomino horses are the result of crosses between horses with black coats and white coats. The alleles for black coats and white coats are codominant.
Which of the following crosses could give palomino offspring?
I. palomino × palomino II. palomino × white III. white × white

Which event happens in meiosis II but not in meiosis I?

Describe the process of crossing over.

Explain the reason for linked genes not following the pattern of inheritance discovered by Mendel.
